
Spatial data is drastically accelerating biological discovery,
changing our perception of biology.
Pharma: Maximize insights from your spatial biology experiments
Our platform integrates spatial multi-omics with histology and single-cell data in one governed analytical layer to unravel complex tissue biology and empower multidisciplinary teams. Give your insights the home they deserve
What Weave delivers
One platform for all stakeholders
Pathology, biology, computational biology, and AI teams work on the same governed dataset. Weave streamlines collaboration through rich visualizations, extensible data analysis pipelines, and deep integration into your existing workflows, across single-cell and multicellular views.
Vendor-neutral end-to-end analysis and integration
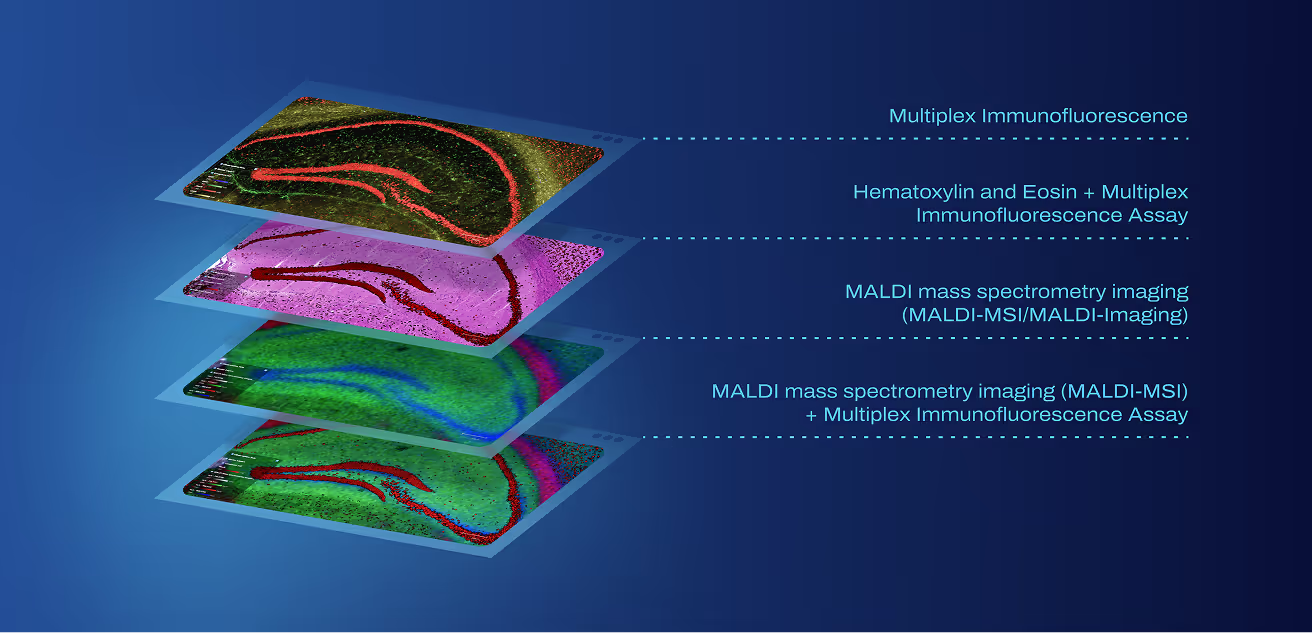
Fuse your spatial transcriptomics, proteomics, metabolomics, mass spectrometry imaging (MSI), histology, and metadata from any source into one analytical asset. Works with your existing tools and infrastructure for digital pathology, LIMS, data catalogs, cloud environment, and computational pipelines.
Trusted insights
Apply QC workflows, provenance, versioning, and model-ready outputs that scale across studies to create high-quality governed datasets with lineage tracking, centralized documentation and flexible access control.
Reusable, governed datasets
Insights compound as cohorts grow and externalize. Deploy results back into your tools via UI or SDK/APIs and control sharing across stakeholders without data duplication.
Proven at
enterprise scale
Supports oncology, immunology, neurodegenerative and other programs
Used from target discovery to translational research
Trusted by the majority of top 10 pharma (including Merck, GSK and BI)
70% faster for integrated results (from 4+ months down to 3 weeks)
Powers studies involving 100s of TB of data and 100M+ cells across multiple modalities
Enables collaboration and data reuse across globally distributed teams
ISO 27001:2022 compliant
"The Aspect Analytics team has a deep understanding of spatial multi-omics data analysis challenges…We had difficulties combining and visualizing results from different spatial-omics modalities…these were solved with user-friendly tools that allow us to maximize information extraction and aid our understanding of mechanism of action for drug molecules."
- Reid Groseclose, Director - Cellular Systems Imaging, GSK
"Leveraging spatial multi-omics in the early phases of drug discovery is a game-changer for accelerating novel target and tissue biomarker efforts. As a pathologist, I really appreciate applications like Weave from Aspect Analytics, which provides intuitive and rigorous tools to explore and integrate spatial transcriptomics and spatial proteomics datasets with traditional pathology imaging. Like a google map microscope, Weave allows us to navigate the tissue multidimensionally and explore spatial relationships at depth. It is helping elevate our understanding of the pathobiology of disease to the next level!"
- Silvia Siso, DVM, MS, PhD, Sr Ppal Scientist-Pathologist, Abbvie

Check out some of our joint publications
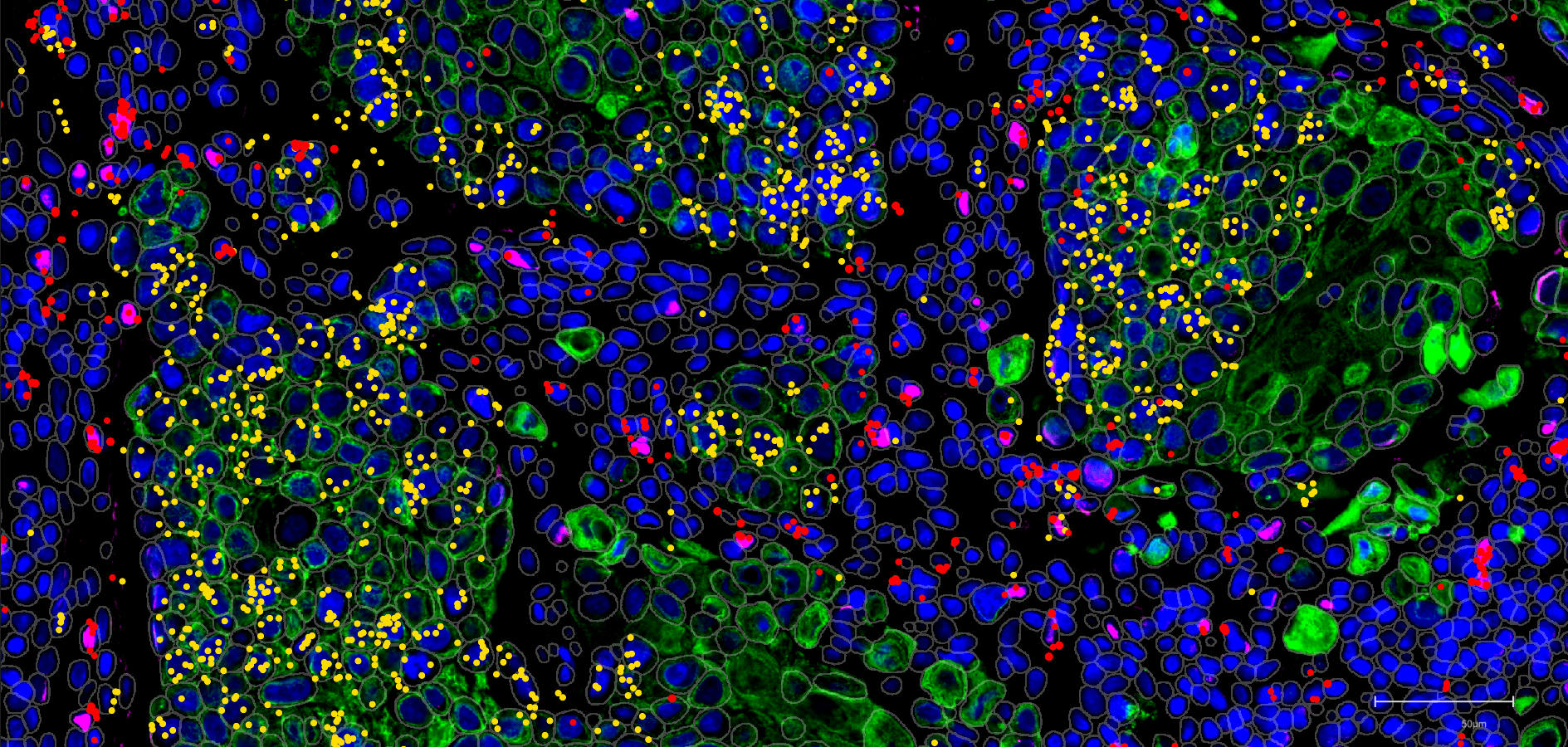

Classification of Small Blue Round Cell Tumors by integrating peptide and N-glycan mass spectrometric profiles

Quantitative MALDI Imaging of Aspirin Metabolites in Mouse Models of Triple-Negative Breast Cancer
Request a demo or run a multicellular cohort analysis with our team
